A molecular simulation environment lets students explore the effect of molecular interactions on the biochemical properties of systems. Biochemistry is an introductory course, designed for both biology and chemical engineering majors.
Learn about Open & Free OLI courses by visiting the “Open & Free features” tab below.
Biochemistry — Open & Free
- Description
- What students will learn
- Learning objectives by module
- Course outline
- Other course details
- System requirements
- Open & Free features
Description
Biochemistry is an introductory course, designed for both biology and chemical engineering majors.
A consistent theme in this course is the development of a quantitative understanding of the interactions of biological molecules from a structural, thermodynamic, and molecular dynamic point of view. A molecular simulation environment provides the opportunity for you to explore the effect of molecular interactions on the biochemical properties of systems.
This course assumes that students have taken introductory chemistry, including basic thermodynamics, as well as introductory organic chemistry. An introductory biology course is not a prerequisite for the course, but students would benefit from some prior exposure to biology, even at the high school level. Required mathematical skills include simple algebra and differential calculus.
In-Depth Description
The two main learning goals of the course are:
- Predicting how changes in structure affect function.
- Utilizing quantitative approaches to characterize structure-function relationships in biochemical systems.
The course begins with amino acids and transitions into protein structure and thermodynamics. Protein-ligand binding is treated for both non-cooperative and cooperative binding using immunoglobulins and oxygen transport as examples. The enzymatic function of proteins is explored using serine and HIV proteases as examples. Enzyme kinetics is treated using steady-state kinetic analysis. Enzyme inhibition is treated quantitatively, using HIV protease as a key example.
Carbohydrate and lipids are presented in sufficient depth to allow the student to fully understand major aspects of central metabolism. The discussion of metabolism is focused on energy generation, fermentation, and metabolic control.
The course concludes with an extensive section on nucleic acid biochemistry. The focus of this section is to provide the student with sufficient background so that they are literate in the recombinant DNA technologies as they relate to protein production using recombinant methods.
After a treatment of molecular forces and solution properties, the course builds on molecular and energetic descriptions of fundamental monomeric building blocks to develop a comprehensive understanding of the biological function of polymers and molecular assemblies at the molecular and cellular level. In addition to multiple case studies, the course concludes with a capstone exercise that leads students through the steps required to produce recombinant proteins for drug discovery. The major topics in the course are:
- Protein function, including oxygen transport, antibody function, and enzyme catalyzed reactions.
- Structure and function of carbohydrates and their importance in central metabolism.
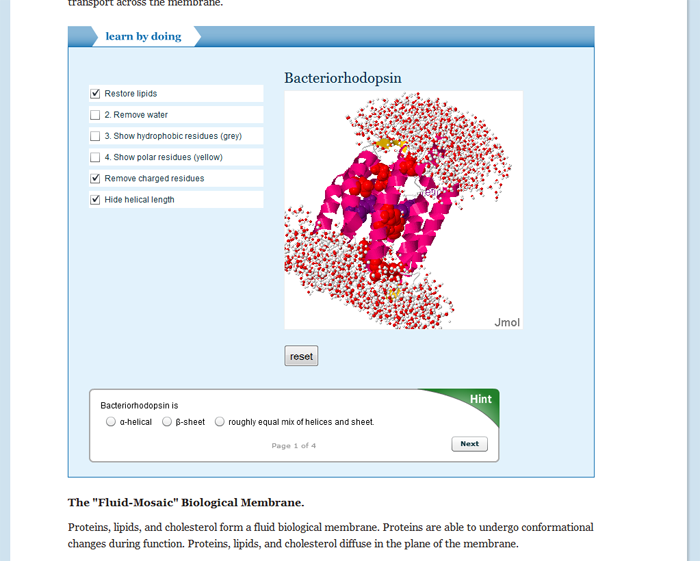
- Lipids and biological membranes.
- Central aspects of metabolism and metabolic control.
- Nucleic acid biochemistry, with emphasis on recombinant DNA technology.
What students will learn
By the time they finish this course, students will learn about or be able to:
- Predict how changes in structure affect function.
- Utilize quantitative approaches to characterize structure-function relationships in biochemical systems.
Learning objectives by module
Unit 2: Biochemistry
Module 2: Introduction and Water Structure
- Characterize functional groups.
- Describe molecular orbitals.
- Explain the molecular structure of Water.
- Identify hydrogen bond donors and acceptors.
- Predict the solubility of compounds in water.
- Predict the strength of hydrogen bonds based on geometry.
- Provide a general overview of biochemistry based on component parts.
- Recognize chiral centers
Module 3: Acids and Buffers
- Calculate the amount of a weak acid and its conjugate base to make a buffer system.
- Calculate the amount of an acid that is protonated at any given pH.
- Calculate the net charge on a molecule, at any pH.
- Understand the relationship between the structure and acidity of an acid.
- Understand why solutions of weak acids resist pH changes.
Module 4: Protein Structure
- Characterize molecular forces between antibody and antigen.
- Define the terms antigen, epitope, and hapten.
- Describe the process of antibody production by B-cells.
- Determine the interaction of each amino acid with water.
- Distinguish between primary, secondary, tertiary and quaternary structure.
- Draw the structure of all amino acids.
- Generate the primary sequence from sequencing data.
- Identify chiral centers on amino acids.
- Interpret changes in ΔH and ΔS.
- Join two amino acids into a dipeptide.
- Obtain ΔH and ΔS for protein unfolding from experimental data.
- Predict degree of protein unfolding given ΔH and ΔS.
- Predict the result of treating a protein with different cleavage reagents.
- Predict the secondary structure of a residue from the Ramachandran plot
- Relate thermodynamic forces to the stability of super-secondary structures.
- Understand conformational entropy.
- Understand consequences of orbital overlap on the configuration of the peptide bond.
- Understand dominant enthalpic forces that stabilize proteins.
- Understand geometrical properties of linear polymers.
- Understand the relationship between primary, tertiary, and quaternary structure of antibodies.
- Understand the role of hydrogen bonds in the stability of secondary structures.
- Understand the role of the hydrophobic effect in protein folding.
- Understand why trans peptide bonds are more stable.
Module 5: Binding
- Calculate amount of oxygen delivered from binding curves.
- Compare and contrast homotropic and heterotropic allosteric effectors.
- Compare and contrast structures and binding curves of myoglobin and hemoglobin.
- Construct Hill plot and obtain KD and Hill coefficient.
- Describe how the amount of ligand bound is affected by the ligand concentration, both at low and high [L].
- Determine the dissociation constant, KD directly from binding data.
- How allosteric activators and inhibitors affect binding.
- Interpret KD in terms of inter-molecular interactions.
- Relate the Hill coefficient to the molecular behavior of the system.
- Understand role of allosteric modulators in oxygen delivery.
- Understand the molecular basis of oxygen transport.
Module 6: Enzymes
- Be able to describe how the substrate concentration affects the rate of enzymatic reactions.
- Be able to obtain kinetic parameters from experimental data
- Calculate the rate enhancement due to transition state stabilization.
- Compare and contrast the structure and mechanism of HIV protease to serine proteases.
- Define steady-state conditions
- Describe the factors that contribute to stabilization of the transition state.
- Determine the dissociation constant for inhibitors.
- Distinguish between enthalpic and entropic stabilization of the transition state.
- Explain the difference between competitive and non-competitive inhibitors.
- Identify steps in viral replication that are suitable for inhibition with drugs.
- Interpret KI values within the context of molecular interactions.
- Understand the basis of substrate specificity.
- Understand the difference between reaction rates and KM and kcat.
- Understand the mechanism of serine proteases.
- Understand the relationship between substrate concentration and the rate of product formation.
Module 7: Protein Purification and Structure Determination
- Calculate and interpret specific activity.
- Determine quaternary structure of a protein.
- Develop a purification scheme.
- Distinguish between molecular weight determination methods.
- Distinguish between the importance of amplitude and phase in X-ray structure determination.
- Understand the principles of separation techniques.
Module 8: Carbohydrates
- Compare and contrast the structure of glycogen and cellulose
- Describe how ring structures of saccharides are formed.
- Describe the structure of bacterial cell walls.
- Draw and name disaccharides.
- Draw the chemical structure of aldoses and ketoses.
- Identify the anomeric carbon in cyclic monosaccharides.
Module 9: Lipids
- Calculate the energetics of membrane-protein interactions.
- Describe the thermodynamics of self-assembly of membranes.
- Draw the structure of Phospholipids.
- Draw the structure of saturated and unsaturated fatty acids.
- Draw the structure of triglycerides.
- Predict membrane permeability of compounds.
- Predict the critical micelle concentration of fatty acids, based on structure.
- Predict the melting temperature of fatty acids, based on structure.
- Predict the melting temperature of model bilayers, based on structure.
Module 10: Metabolism
- Be able to describe how intermediates in the TCA cycle are used to synthesize a wide range of compounds.
- Be able to describe how sugar and fatty acid oxidation are connected to the TCA cycle.
- Be able to describe where, and in what form, the energy is stored that is released by the TCA cycle.
- Describe a linear, branched, and circular pathway.
- Describe the common features of metabolic pathways.
- Enzyme Regulation: Role of protein kinases and phosphatases in control of enzyme function.
- Hormonal Control: Understand physiological role of hormones epinephrine, glucagon, and insulin.
- How the energy stored in a thioester can be used for ATP synthesis or organic addition reactions.
- Identify that glycolysis oxidizes the six carbon glucose to produce two three-carbon pyruvate molecules.
- Identify the cellular location of glycolysis.
- Identify the role of direct and indirect coupling in metabolic pathways.
- Know that electron transport flows through complexes I, III, and IV leads to generation of a hydrogen ion gradient across the inner mitochondrial membrane.
- Know the pathway of the electrons from FADH2 to oxygen, resulting in the formation of water.
- Know the pathway of the electrons from NADH to oxygen, resulting in the formation of water.
- Know the role of the four complexes, coenzyme Q, and cytochrome C in electron transport.
- Predict the direction of reactions using Gibbs free energy.
- Predict the product of oxidative decarboxylations
- Signal transduction: Understand the role of G-proteins and adenyl cyclase in receptor mediated signal transduction.
- State the major differences between catabolic and anabolic pathways.
- Understand activation of enzymes by allosteric compounds as a method of regulation.
- Understand anaerobic metabolism and its role in allowing glycolysis to continue in the absence of oxygen.
- Understand coordinated regulation of opposing steps in synthetic and degradative pathways.
- Understand glucose can be synthesized from pyruvate.
- Understand how both direct and indirect coupling are used to make glycolysis spontaneous.
- Understand how glycogen degradation and synthesis is controlled by protein phosphorylation.
- Understand how the energy released by the oxidation is stored as ATP and “high-energy” electrons.
- Understand regulation by non-covalent and covalent modification of enzymes.
- Understand that the free energy stored in the non-equilibrium hydrogen ion gradient is sufficient to sythesize ATP from the transport of 3 protons.
- Understand the major differences between a competitive and uncompetitive inhibitor.
- Understand the major energy storage methods in metabolism – phosphorylated compounds, redox carriers, proton gradient.
- Understand the mechanism of ATP synthase, how allosteric changes that occur during proton translocation lead to the conversion of ADP and Pi to ATP.
- Understand the relationship between the Gibbs free energy change of a step in the pathway and whether the step can be accomplished by the same enzyme in the reverse direction.
- Understand the source of “high-energy” phosphate bonds in ATP
- Understand the structure of glycogen.
Module 11: Recombinant DNA and Central Dogma
- Be able to utilize the lac repressor/operator to control gene expression
- Calculate TM of a double stranded DNA molecule.
- Compare and contrast the major factors that stabilize dsDNA and Proteins.
- Describe the overall process of amino acid synthesis on the ribosome
- Describe the role of tRNA structure in protein synthesis.
- Design a coding sequence with the correct signals for protein synthesis
- Design primers for PCR DNA amplification.
- Determine a protein sequence given a sequencing gel.
- Discuss the major features that affect the energetics of DNA-protein interactions.
- Discuss the role of each element in an expression vector.
- Distinguish between a purine and a pyrimidine.
- Draw the monomeric structure of a nucleotide.
- Explain how a polymer of nucleic acid is formed from nucleoside triphosphates.
- Explain how PCR leads to the amplification of DNA.
- Explain how proofreading occurs during DNA synthesis.
- Explain the basis of DNA sequencing.
- Explain the difference in the chemical stability of RNA versus DNA.
- Explain the effect of salt concentration on the melting temperature of DNA.
- Explain why DNA synthesis is always in the 5′-3′ direction.
- Explain why there is a high mutation rate in the HIV virus.
- Identify the major and minor group of a base pair.
- Identify the non-Watson-Crick hydrogen bonds in A-T and G-C basepairs.
- Identify the Watson-Crick hydrogen bonds in A-T and G-C basepairs.
- Include the appropriate signals to cause export of the protein out of the cell
- List the essential elements of an expression vector.
- Provide a description of the structural features of B-DNA.
- The describe the mechanism of RNA polymerase
- Use restriction enzymes and DNA ligase to generate recombinant DNA molecules.
Module 12: DNA Replication
- Describe the activities of each protein involved in replication.
- Understand the enzymatic differences in polymerases and the importance of these differences in functionality
Course outline
UNIT 1: Overview
Module 1: Spring 2017
UNIT 2: Biochemistry
Module 2: Introduction and Water Structure
Module 3: Acids and Buffers
Module 4: Protein Structure
Module 5: Binding
Module 6: Enzymes
Module 7: Protein Purification and Structure Determination
Module 8: Carbohydrates
Module 9: Lipids
Module 10: Metabolism
Module 11: Recombinant DNA and Central Dogma
Module 12: DNA Replication
UNIT 3: Glossaries
Module 13: Glossaries
Other course details
System requirements
OLI system requirements, regardless of course:
- internet access
- an operating system that supports the latest browser update
- the latest browser update (Chrome recommended; Firefox, Safari supported; Edge and Internet Explorer are supported but not recommended)
- pop-ups enabled
- cookies enabled
Some courses include exercises with exceptions to these requirements, such as technology that cannot be used on mobile devices.
This course’s system requirements:
Open & Free features
Open & Free Courses
- Open & Free OLI courses enable independent learners to study a subject on their own terms, at their leisure. Courses are:
- Self-guided.
- Self-paced.
- Self-supported.
- Open & Free courses include only the learning materials:
- No teacher.
- No tests.
- No college credit.
- No certificate of completion.
- *If your teacher gave you a Course Key, do not use an Open & Free course because your teacher will never see your work.